kMoL, a Machine Learning Library for AI Drug Discovery With Federated Learning Capabilities
Elix, Inc., an AI drug discovery company with the mission of “Rethinking Drug Discovery” (CEO: Shinya Yuki/Headquarters: Tokyo, Japan; hereinafter referred to as “Elix”) has developed kMoL, a machine learning library for AI drug discovery with federated learning functionally. This work has been discussed and developed with lecturer Ryosuke Kojima and Professor Yasushi Okuno from the Graduate School of Medicine, Kyoto University. It has since been released as open-source on October 20th, 2021.
kMoL is a library for building machine learning models for the drug discovery and life science fields. This library has been improved based on the knowledge obtained from kGCN(1), an open-source AI library for drug discovery and the life sciences that has been developed by lecturer Ryosuke Kojima and Professor Yasushi Okuno from the Graduate School of Medicine, Kyoto University. It also includes graph neural networks that can handle graph structures that are widely useful in life sciences, such as molecular structures and chemical pathways.
One of the most significant features of kMoL is that it is the only publicly available library for AI drug discovery that has a “federated learning” function. As federated learning enables the library to access huge amounts of data while ensuring security, it has recently attracted attention as a learning method for handling confidential information, such as compound data, in the pharmaceutical industry. Included as a part of kMoL is Elix’s federated learning library, Elix Mila.
As kMoL supports advanced models with a wide range of applications and can securely access vast amounts of data for learning, it is expected to be widely adopted by pharmaceutical and chemical companies.
Overview of kMoL: a machine learning library for AI drug discovery with federated learning

Name: kMoL (Machine Learning library for Molecular systems)
Abstract: A machine learning library for AI drug discovery with federated learning capabilities. It has features such as support for federated learning and graph-based predictive models.
Release date: October 20, 2021
Open-source URL: https://github.com/elix-tech/kmol
This library has been developed in collaboration with Elix and lecturer Ryosuke Kojima and Professor Yasushi Okuno under a contract from Kyoto University, which has executed a research consignment contract with the Japan Agency for Medical Research and Development (AMED) under the “Development of a Next-generation Drug Discovery” (DAIIA).
Functions and features of kMoL
1.Support for federated learning
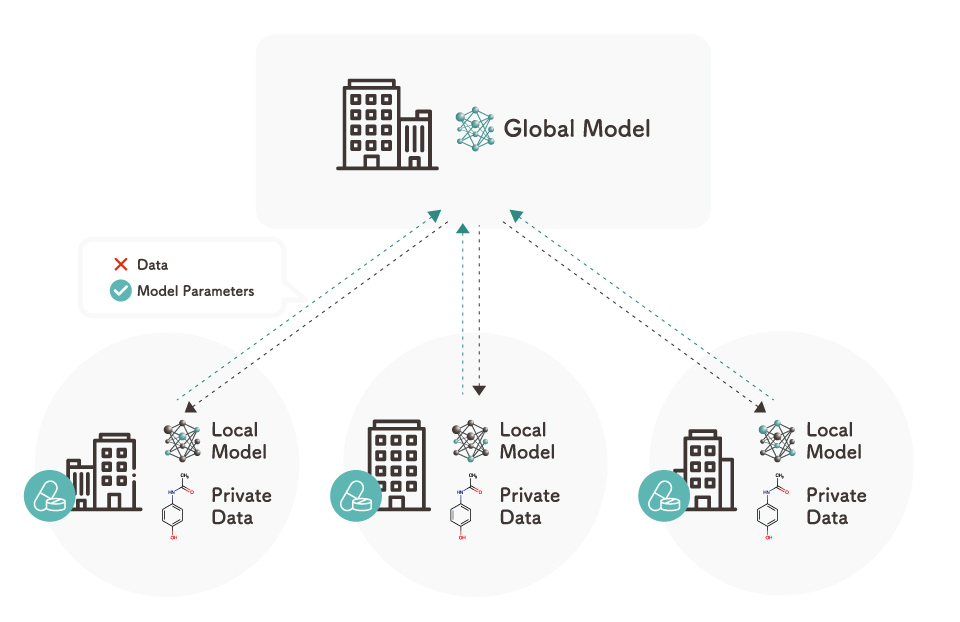
Federated learning is a method of machine learning in which data is not aggregated but is instead distributed (i.e., data is not shared outside the company). In industries that handle highly confidential data, it is difficult to share data. Therefore, federated learning is gaining attention as a method to ensure data privacy and security.
kMoL incorporates Elix Mila, a federated learning module developed by Elix, making kMoL the only machine learning library with federated learning capabilities among those released for AI drug discovery. Using this library makes it possible to utilize a large amount of data for learning without compromising the confidentiality of compound data. As machine learning models often benefit from large amounts of data, federated learning enables more data to be used to attain higher accuracy without compromising the confidentiality of compound data.
[Diagram of federated learning]
2.Support for graph-based predictive models
One of the best features of kMoL as a machine learning library for life sciences is that it can seamlessly use state-of-the-art graph-based predictive models with federated learning. One of the best ways to represent a molecule’s structure is by using a graph. As a result, graph-based predictive models have a significant advantage over other architectures as they can utilize far more information about the molecular structure of a compound. This extra information is expected to greatly increase the accuracy of learning.
kMoL has been validated on ADME (A: absorption, D: distribution, M: metabolism, E: excretion), toxicity and binding affinity datasets. It is also possible to learn and predict unique tasks for unique datasets.
3.Other features
Another feature of kMoL is that it can be used with the machine learning framework PyTorch. When Elix started developing kMoL, most machine learning libraries with federated learning capabilities were based on the machine learning framework TensorFlow. PyTorch is currently one of the most popular machine learning frameworks owing to the ease of model implementation(2) and, to make it available to a wider audience, kMoL supports model development based on PyTorch.
In addition, to protect data privacy, some models also support a technique called differential privacy. This is a method that makes it impossible to distinguish as to which data contributed to the model while minimizing the impact on prediction accuracy. kMoL can also run on both GPUs and CPUs, a feature that was not supported by the previously released machine learning libraries with federated learning capabilities.
Shinya Yuki, CEO of Elix, Inc. said: “We are very pleased to release kMoL as an open-source library that we have jointly built based on our Elix Mila federated learning module and Kyoto University’s kGCN. By combining federated learning with predictive models, we will be able to achieve things that cannot be done by a single organization. We hope that this library will accelerate drug discovery research and contribute to the development of this field.”
Professor Yasushi Okuno, Graduate School of Medicine, Kyoto University said: “The remarkable progress of AI in recent years has had a powerful impact on drug development. Led by Lecturer Kojima, we have been developing world-leading AI programs for drug discovery, by developing deep learning technologies to deal with the chemical structures of drugs and molecular networks in living organisms. We are now collaborating with Elix to develop a federated learning package in addition to the technologies we have developed so far and release it as the drug discovery AI library ‘kMoL.’ We hope that this library will be widely applied in industry through Elix.”
He also added “kMoL is an extension of the drug discovery AI library ‘kGCN’ that has been developed by the research team of lecturer Ryosuke Kojima and Professor Yasushi Okuno. The federated learning function of this software was developed as part of the ‘Development of a comprehensive drug discovery AI platform combining multi-target prediction and structure generation using state-of-the-art AI technology’ project under the ‘Drug Discovery Support Promotion Project: Development of the Next Generation Drug Discovery AI through the Industry-Academia Collaboration (DAIIA)’ of the Japan Agency for Medical Research and Development (AMED).
In addition, the multimodal neural network incorporates the knowledge accumulated through the results of the ‘Development of AI for drug formulation design to improve efficiency and accelerate drug development’ project, organized by the New Energy and Industrial Technology Development Organization (NEDO).
The large-scale graph neural network incorporates the knowledge acquired through the results of the ‘Construction and expansion of case database and development of drug target estimation algorithm to accelerate new drug creation’ project, organized by the Public-Private R&D Investment Strategic Expansion Program PRISM.”
Reference
1. R. Kojima, S. Ishida, M. Ohta, H. Iwata, T. Honma, Y. Okuno: kGCN: a graph-based deep learning framework for chemical structures. Cheminformatics, Springer, Vol. 12, pp. 1–10, 2020.
2. From The Gradient presentation “The State of Machine Learning Frameworks in 2019” (October 2019). https://thegradient.pub/state-of-ml-frameworks-2019-pytorch-dominates-research-tensorflow-dominates-industry/
The most recent data can be viewed at http://horace.io/pytorch-vs-tensorflow/.
About Elix, Inc.
Elix, Inc is an AI drug discovery company with the mission of “Rethinking drug discovery”. To significantly improve the time-consuming and expensive drug discovery process, we have applied cutting-edge deep learning and machine learning technologies to develop business for a variety of clients. These include pharmaceutical companies, chemical companies, and universities.
Visit https://www.elix-inc.com/ for more details.
About lecturer Ryosuke Kojima and Professor Yasushi Okuno from the Graduate School of Medicine, Kyoto University
Lecturer Ryosuke Kojima and Professor Yasushi Okuno from the Graduate School of Medicine, Kyoto University, aim to pioneer simulation science and data science for medical and drug discovery applications. They develop new methodologies for medical big data analysis and medical simulation using actual clinical data from Kyoto University Hospital and work on drug discovery simulation and big data drug discovery using the supercomputer Fugaku to achieve their goal.
Visit http://clinfo.med.kyoto-u.ac.jp/en/ for more details
